Posted: March 30, 2023
Novel approaches to food safety are rooted in the detection of foodborne pathogens.
One of the most important missions of Penn State's College of Agricultural Sciences always has been to make the food supply safer. Today, a cross-disciplinary team of faculty is conducting research to leverage high-technology approaches to reduce the incidence of foodborne illness.
That mission is critical. In the U.S. alone, the Centers for Disease Control and Prevention estimates that roughly one in six Americans -- or 48 million people -- get sick each year; 128,000 are hospitalized, and 3,000 die of foodborne diseases.
Foodborne diseases are caused by food contamination and occur at any stage of the food production, delivery and consumption chain. They can result from several forms of environmental contamination, including pollution in water, soil or air, as well as unsafe food storage and processing. Hundreds of diseases, according to the CDC, are caused by eating food contaminated with bacteria, viruses, parasites or chemical substances.
"Our researchers, in many cases working with state and federal agencies, are developing innovative technologies and surveillance systems to make the food system safer," said Blair Siegfried, associate dean for research and graduate education in the college and director of the Pennsylvania Agricultural Experiment Station. "Our faculty, several of whom are just launching their careers, are employing cutting-edge technologies to minimize negative impacts on public health. Projects include gene tracing, post-microbe interactions, preventing spoilage and food waste, and microbe interactions for food safety and animal production efficiency."
Here is a sampling of work in progress.
Peeling Back the Mystery of Biofilms
Last year, microbiologists in the college received a $605,000 grant from the U.S. Department of Agriculture to study how microbial biofilms protect Listeria monocytogenes, the bacterium that causes the deadly foodborne illness listeriosis.
Jasna Kovac, the Lester Earl and Veronica Casida Career Development Professor of Food Safety, and Luke LaBorde, professor of food science, are using the funding from USDA's National Institute of Food and Agriculture to conduct research on the interactions between microorganisms found in fruit-packing environments and Listeria monocytogenes.
"We are studying the ability of environmental microorganisms to form robust biofilms together with L. monocytogenes and how these biofilms may protect L. monocytogenes from the antimicrobial activity of sanitizers," said Kovac, assistant professor of food science. "The data generated in this project will help improve the cleaning and sanitizing used in the fresh produce industry to better control L. monocytogenes and support safe food production."
Listeria and other microorganisms found in the natural environment, such as soil, can be introduced unintentionally into facilities that process raw foods such as fruit. The research is needed, Kovac explained, because once introduced into the food-processing environment, Listeria and many other environmental microorganisms can grow on surfaces into microbial layers called biofilms.
"Microorganisms enclosed in a biofilm produce slimy substances that protect them from the antimicrobial activity of sanitizing chemicals by slowing down their penetration into the core of a biofilm," Kovac said. "Biofilm formation can therefore result in reduced efficacy of antimicrobial sanitizers used to inactivate Listeria."
Graduate student Laura Rolon is working to isolate environmental microbiota and determine their resistance to sanitizers, characterizing cultures of microorganisms recovered from food-processing facilities using whole-genome sequencing. "Listeria monocytogenes is especially dangerous because the pathogen can survive, grow and persist at low temperatures in produce-processing facilities," she said.
Researchers and industry still do not have a clear understanding of the ways biofilms inhibit sanitization within the food-processing environment, LaBorde explained. So, he and his fellow researchers will investigate how environmental microbiota found in produce-packing environments form single- and multi-species biofilms with L. monocytogenes.
GenomeTrakr Network
Scientists who track outbreaks of foodborne illness are left to ask questions such as, "Was it the lettuce in the salad, the meat on the sandwich or the fresh fruit cup that caused 25 people from a restaurant to become violently ill? Did a virus or bacterium sneak in on the hands of an employee or hitchhike on food all the way from the farm?" Tracing the source of an outbreak historically has taken months. But time is imperative when people are getting sick and even dying.
For that reason, in 2016, Penn State became one of the first academic institutions to take the lead for its state in the U.S. Food and Drug Administration's GenomeTrakr network. Started in 2012, GenomeTrakr is a nationwide system of laboratories that utilize whole-genome sequencing -- which identifies the entire genetic blueprint for particular microbial species -- to rapidly and conclusively identify pathogens involved in outbreaks of foodborne illness.
Ed Dudley, professor of food science, who is spearheading Pennsylvania's work with GenomeTrakr, explains why Penn State got involved.
"Our goal is to help populate the whole-genome-sequence databases with foodborne isolates, particularly E. coli and Salmonella, from food and environmental sources," he said.
For years, Dudley and the students working in his laboratory have conducted research using molecular biology and biochemistry to understand better the physiology, behavior and evolution of foodborne pathogens with an eye toward improving methods of tracking the spread of the organisms "from farm to fork."
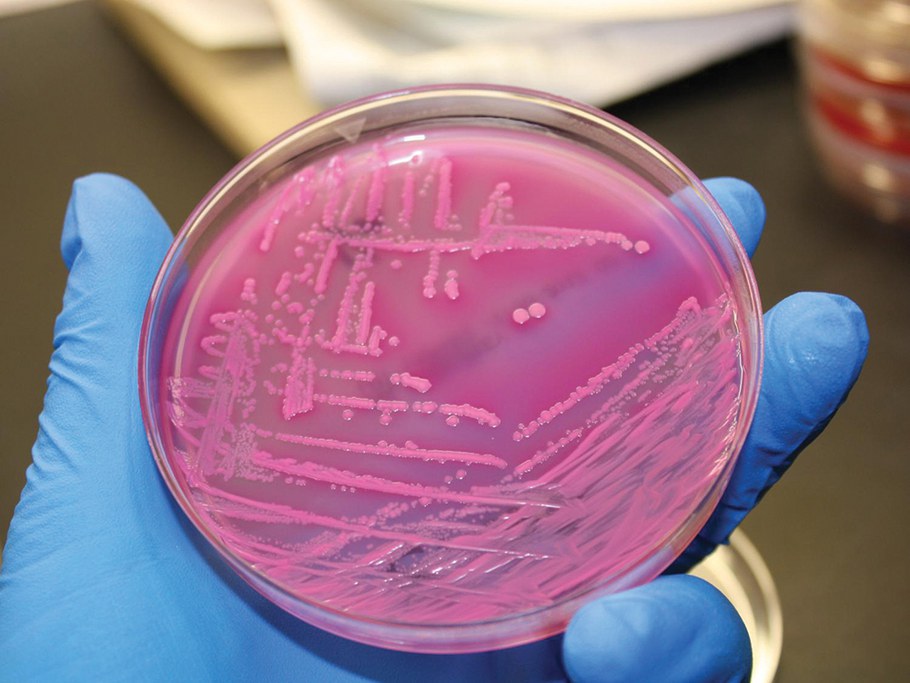
E. coli, shown here in an agar plate, is the best studied bacterium on the planet. It is often responsible for outbreaks of foodborne illness.
The GenomeTrakr network currently consists of 15 federal labs, 25 state health and university labs, one U.S. hospital lab, two other labs located in the U.S., and 20 labs outside of the U.S. They collect and share genomic and geographic data from foodborne pathogens.
Researchers and public health officials can access the data -- which are housed in public databases at the National Center for Biotechnology Information at the National Institutes of Health -- for real-time comparison and analysis. Easy access to that vital information will aid investigations on foodborne illness outbreaks and reduce illnesses and deaths.
Banking Bacteria
According to Dudley, the college's E. coli Reference Center, housed in the Department of Food Science, was one reason Penn State was selected for inclusion in the GenomeTrakr network.
"It's the world's largest E. coli collection, with 95,000 isolates from food and animals collected over 60 years, that provides us with a rich source of bacteria to sequence," he said.
By analyzing the whole genomes of these bacteria, Dudley and his team will add significantly to the FDA's databases, which, in turn, will help the organization to track the sources of even more outbreaks.
Recently Dudley and the college received a four-year, $371,000 grant from the National Science Foundation to be part of a multi-institutional, $2.5 million research project aimed at predicting bacteriophage resistance from only a genome sequence. With more than 6,200 of its sample genomes sequenced, the E. coli Reference Center is likely the only publicly available collection in the country that could have been leveraged for this particular project, according to Dudley.
"We are using bacterial genome sequences to predict bacteria sensitivity/resistance to bacteriophage," he said. "This is relevant to phage therapy as it could help speed selection of bacteriophage capable of killing a specific strain isolated from a patient."
A bacteriophage is a type of virus that infects only bacteria, Dudley said. In fact, the word "bacteriophage" literally means "bacteria eater," because bacteriophages destroy their host cells. A bacteriophage attaches itself to a susceptible bacterium and infects the host cell. Following infection, the bacteriophage hijacks the bacterium's cellular machinery to prevent it from producing bacterial components and instead forces the cell to produce viral components.
So-called phage therapy is seen as a possible weapon against multi-drug-resistant strains of many bacteria, Dudley said.
Finding the Canary in the Mine
In a completely different approach, Erika Ganda, assistant professor of food animal microbiomes, is using machine-learning algorithms to detect abnormalities in microbiome data in food-production systems. She believes this revolutionary method, made possible using supercomputers and artificial intelligence for anomaly detection, is an important next step toward bettering our food supply chain. This project is a partnership with IBM research and leverages the unparalleled expertise of IBM in advanced analytics with Penn State's expertise in microbiomes in the food supply.
Looking for contaminants in raw materials is difficult, she pointed out, and until recently scientists had to test for pathogens individually, one sample at a time. That means running hundreds, if not thousands of tests, most of which are negative. Now, thanks to Ganda and her team, scientists can monitor all the harmless bacteria found in normal, safe food. If those baseline bacteria suddenly change, it could signal contamination.
"Every product has microbes, and we are able to watch these microbes over time in the food-processing facility," Ganda said. "And then if they suddenly change, that gives us a quick clue to ask, 'Hey, what's going on here?' That's the canary in the mine that would allow us to pick out problems earlier."
In an ongoing study, focused on detecting anomalous content in dairy with whole metagenome sequencing, Ganda is leading a research team that has demonstrated that the concept is valid in fluid milk as a model system.
Artificial intelligence was able to classify abnormal versus baseline samples and flag spikes in microbial populations such as Morganella, Enterobacter and Coxiella in association with antibiotic use, and distinguish microbial signatures related to milk collected from alternative farms, the researchers reported. This work also characterized the microbiome of fluid milk in greater sequencing depth than previously published studies.
"Our results indicate the application of artificial intelligence continues to hold promise in the realm of microbiome data analysis and could present further opportunity for downstream analytic automation to aid in food safety and quality," said Ganda. "We evaluated the feasibility of using untargeted metagenomic sequencing of raw milk for detecting anomalous food-ingredient content with artificial intelligence methods. The approach could potentially be applied in the food industry for safety and quality control."
Food scientists can easily distinguish between bacteria that don't cause disease, those that do, and others that are normally found beside or partnering with the pathogens, Ganda explained.
"Sometimes, even though you don't find the disease-causing bacteria, you find its partner, and that could be very useful from a food-safety perspective because then you'll know you need to keep looking," she said. "It's not a diagnostic method, rather it's a kind of a survey or monitoring."
'Huck Hits' Grant
In a separate project, Ganda recently received funding from Penn State's Huck Institutes of the Life Sciences to support her research on real-time and comprehensive antimicrobial-resistance profiling. The study responds to the global threat posed by antimicrobial resistance, which is projected to account for 10 million deaths annually by 2050.
Current methods for tracking resistome -- or the presence of antibiotic-resistance genes -- are labor-intensive or prohibitively expensive for broad surveillance purposes. Ganda's team developed a real-time resistome-profiling system combining several cutting-edge technologies that had never been applied to investigate antimicrobial resistance. Greatly simplified, the method would enrich antimicrobial-resistance genes and sequence them using a pocket-sized sequencer.
"If successful, this method -- the subject of a provisional patent application -- will revolutionize the fight against antimicrobial resistance," Ganda said. "This is a new approach that has not previously been attempted."
Frustrated by Fungi
Bacterial pathogens are not the only threat to the food supply. Mycotoxins produced by fungi pose a significant menace to food safety and nutrition security by contaminating food and feed crops. Existing mitigation strategies for control are inadequate, according to a team of Penn State researchers led by Josephine Wee, assistant professor of food science.
"The mycotoxin problem is really old -- at least 100 years old," Wee said. "There needs to be a renewed research focus on it. In the past, we studied one toxin produced by one fungus. Now the field is moving toward simultaneously looking for many toxins produced by multiple fungi, more of a mycotoxin profile kind of surveillance."
The team, which includes Seogchan Kang, professor of plant pathology and environmental microbiology, and Joshua Kellogg, assistant professor of metabolomics, working out of the Huck Institutes of the Life Sciences, proposes to harness the vast metabolic diversity created via microbial evolution and chemical ecology to provide a safe and cost-effective strategy to diminish mycotoxin contamination.
The researchers have initiated an effort to identify specific bacteria and fungi that can suppress growth of toxigenic Aspergillus mold (a type of fungus) and its production of aflatoxin, the most potent naturally occurring carcinogen.
Members of the fungal family Aspergillus, Penicillium and Fusarium secrete mycotoxins that contaminate staple food and feed crops during cultivation and storage. The U.N. Food and Agriculture Organization estimates that 25% of the world's food crops are contaminated with mycotoxins, causing a massive burden on global food safety and nutrition security. Five billion people, including many in developing countries, are frequently exposed to mycotoxins through their diets and via inhalation.
However, compared to foodborne bacteria, the impact of mycotoxins has received limited attention, Wee pointed out. One reason may be that mycotoxins do not cause acute symptoms unless ingested in large quantities, making them a stealthy and chronic threat.
"In developing countries, this threat is further compounded by poorly established monitoring systems for threats to food and feed safety," Wee said. "But aflatoxin is a big concern in the U.S. when it comes to the export of grains. When the grains reach trade borders, they must meet the limits set forth by the importer or be turned away. For example, the EU has 10 times lower limits for aflatoxin than the U.S."
The removal of mycotoxins from contaminated crops in the field is impractical and not cost effective, the researchers say, so current mitigation strategies focus on preventing contamination during the pre- and postharvest stages.
"However, available strategies are limited, and unreliable," Wee says. "The long-term goal of the research is to significantly reduce mycotoxin burden on animals and humans by developing effective and sustainable strategies based on evolution-driven chemical ecology."
She explained that the researchers will identify potential metabolite regulators produced by fungi and bacteria that reduce aflatoxin and other mycotoxins on food security crops.
Early Detection of Novel Diseases
Sometimes in the Department of Food Science, genomic research to benefit food safety bleeds into studies aimed at enhancing the early detection of novel infectious bacteria that could cause outbreaks of infectious disease and public health emergencies. In one such project, a team of researchers in the college is sequencing the genomes of 700 Bacilli bacteria -- near relatives of the biothreat pathogen that causes anthrax.
Funded by a $1.2 million grant from the U.S. Centers for Disease Control and Prevention, the research will support the development of genomic resources and DNA sequence databases for the federal agency to increase its capacity to rapidly detect novel pathogens, according to team leader Kovac.
"You may have heard of the 2001 bioterrorist attacks in which spores of the bacteria Bacillus anthracis that cause anthrax were circulated in the mail," she said. "People who inhale these spores can get sick with anthrax, which is often fatal."
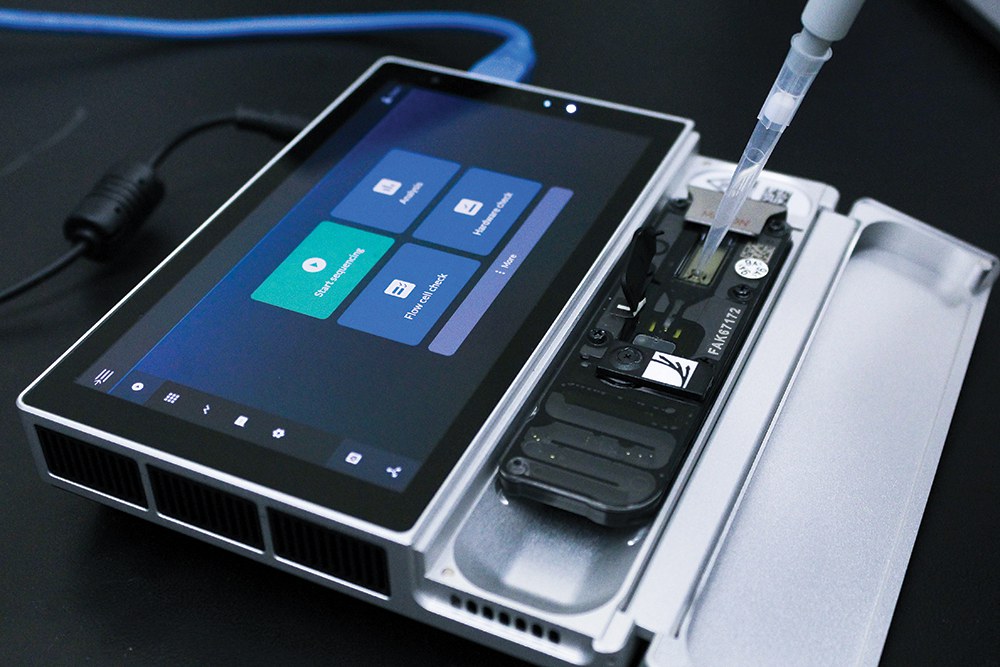

A nanopore sequencer will be used in the study to sequence the genomes of 700 Bacilli bacteria -- near relatives of the biothreat pathogen Bacillus anthracis.
From a biodefense standpoint, it is important to understand the diversity of environmental Bacilli that could become novel biothreats such as anthrax, added Kovac, who has extensive experience with the genomics of Bacilli.
"There are known examples among Bacillus cereus group of bacteria where 'benign' environmental strains have acquired anthrax-causing capabilities," she said. "We are interested in detecting and characterizing similar strains of Bacilli that have both the characteristics of known biothreats and harmless environmental microorganisms."
If emerging pathogens or biothreats are detected early on, they are more likely to be contained effectively to prevent a public health emergency, Kovac noted.
"We are partnering with the CDC to create a large database of Bacilli to support its development of rapid laboratory methods for the detection of novel, naturally occurring or engineered pathogens and potential emerging biothreats," she said.
Penn State researchers are uniquely positioned to complete the proposed work and support the CDC's expansion of reference databases for the detection of novel, emerging infectious diseases, Kovac contends.
"Here in the Department of Food Science, we have microbiology and genomic expertise and access to a large number of unique, environmental and food Bacilli, deposited in the Food Microbe Tracker culture collection," she said.
Strengthening Infectious Disease Surveillance
A recent collaboration with the U.S. Centers for Disease Control and Prevention is another example of using expertise gained in the pursuit of molecular food science for national defense. The federal agency provided a $750,000 grant to fund research by a team of Penn State scientists to strengthen infectious disease surveillance, detection and preparedness by developing an accessible bioinformatics platform and tools for utilization by the CDC's Laboratory Response Network.
The Laboratory Response Network is a linked system of labs across the country that rapidly responds to biological and chemical threats and public health emergencies. The Penn State team will provide the network with an accessible software platform and tools for whole-genome sequencing data analyses and interpretation of results, according to team principal investigator Kovac.
Genomics approaches can be leveraged to detect known and novel infectious agents, including emerging infectious disease pathogens, explained Kovac.
"In this project, our team is partnering with the CDC to assess machine-learning-based approaches for the detection of potential novel pathogens," she said. "We will evaluate the performance of multiple machine-learning-based pathogen-prediction models. The performance of these models will be assessed on a set of Bacilli genomes that we whole-genome sequenced over the past year."
Under the leadership of team co-principal investigator Greg Von Kuster, Penn State Galaxy programming consultant, working out of the Huck Institutes of the Life Sciences, the team will develop a computer-based environment using the platform Galaxy to enable CDC Laboratory Response Network scientists to conduct near real-time bioinformatic analyses and produce actionable information.
By Jeff Mulhollem
Features
Fostering Forests
Across the United States, forests face unprecedented threats, and scientists in Penn State's College of Agricultural Sciences are conducting novel and complex research to conserve them.
Buzzing With Purpose
Community scientists work to protect Pennsylvania's wild bees
Conservation Reimagined
Exploring new approaches to cope with a changing climate